Note
Click here to download the full example code
MDF-based DOE on the Sobieski SSBJ test case¶
from __future__ import division, unicode_literals
from os import name as os_name
from gemseo.api import configure_logger, create_discipline, create_scenario
from gemseo.problems.sobieski.core import SobieskiProblem
IS_NT = os_name == "nt"
configure_logger()
Out:
<RootLogger root (INFO)>
Instantiate the disciplines¶
First, we instantiate the four disciplines of the use case:
SobieskiPropulsion,
SobieskiAerodynamics,
SobieskiMission
and SobieskiStructure.
disciplines = create_discipline(
[
"SobieskiPropulsion",
"SobieskiAerodynamics",
"SobieskiMission",
"SobieskiStructure",
]
)
Build, execute and post-process the scenario¶
Then, we build the scenario which links the disciplines
with the formulation and the optimization algorithm. Here, we use the
BiLevel formulation. We tell the scenario to minimize -y_4
instead of minimizing y_4 (range), which is the default option.
We need to define the design space.
design_space = SobieskiProblem().read_design_space()
Instantiate the scenario¶
scenario = create_scenario(
disciplines,
formulation="MDF",
objective_name="y_4",
design_space=design_space,
maximize_objective=True,
scenario_type="DOE",
)
Set the design constraints¶
for constraint in ["g_1", "g_2", "g_3"]:
scenario.add_constraint(constraint, "ineq")
Execute the scenario¶
Use provided analytic derivatives
scenario.set_differentiation_method("user")
n_processes = 4
if IS_NT: # Under windows, don't do multiprocessing
n_processes = 1
We define the algorithm options. Here the criterion = center option of pyDOE centers the points within the sampling intervals.
algo_options = {
"criterion": "center",
# Evaluate gradient of the MDA
# with coupled adjoint
"eval_jac": True,
# Run in parallel on 4 processors
"n_processes": n_processes,
}
run_inputs = {"n_samples": 30, "algo": "lhs", "algo_options": algo_options}
scenario.execute(run_inputs)
Out:
{'eval_jac': False, 'algo': 'lhs', 'n_samples': 30, 'algo_options': {'criterion': 'center', 'eval_jac': True, 'n_processes': 4}}
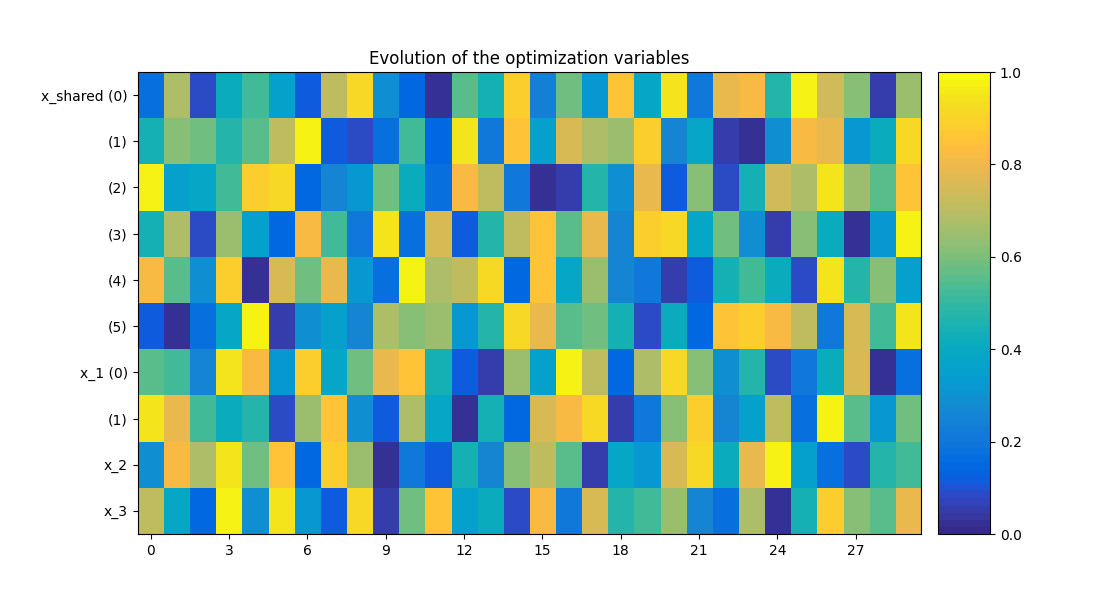
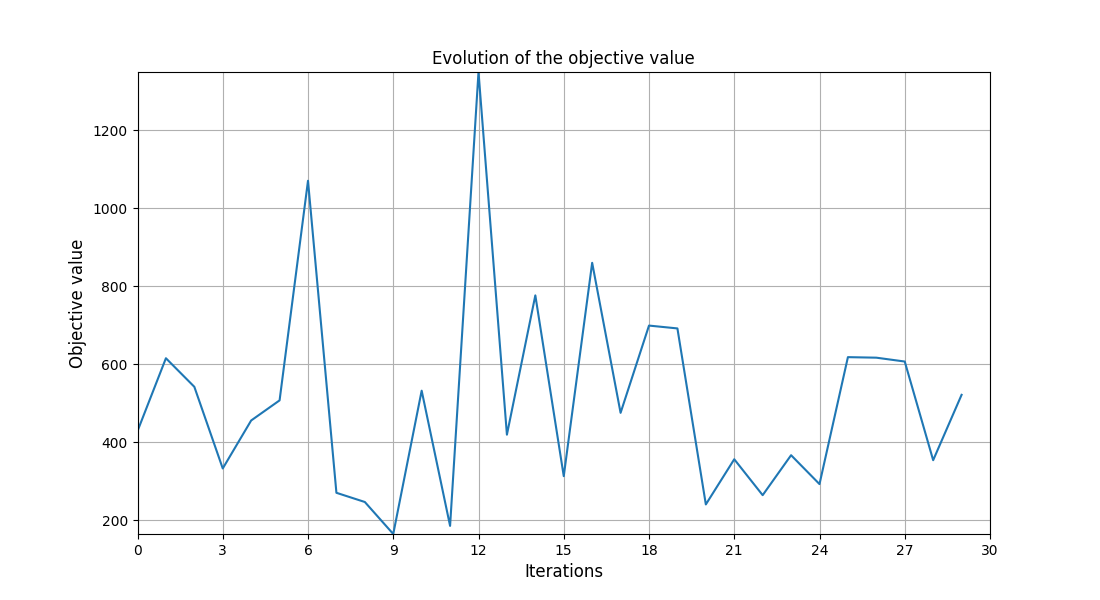
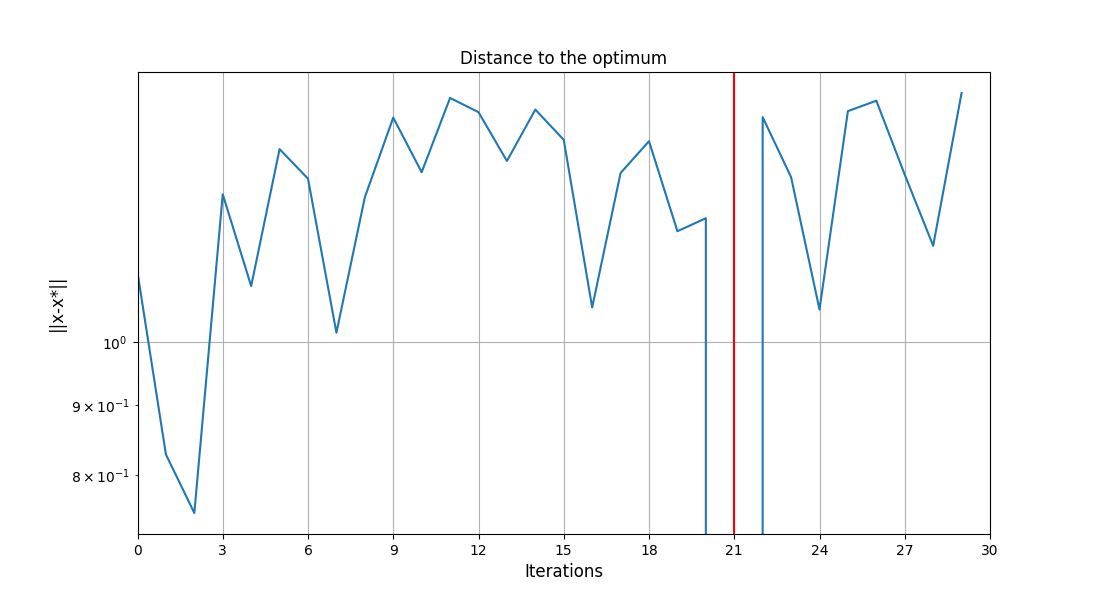
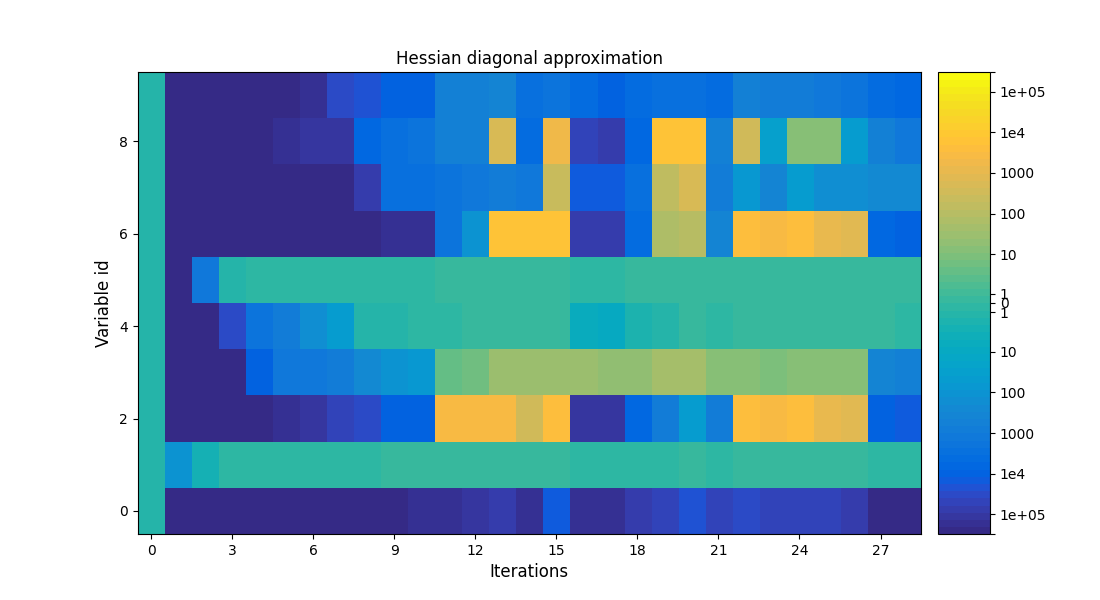
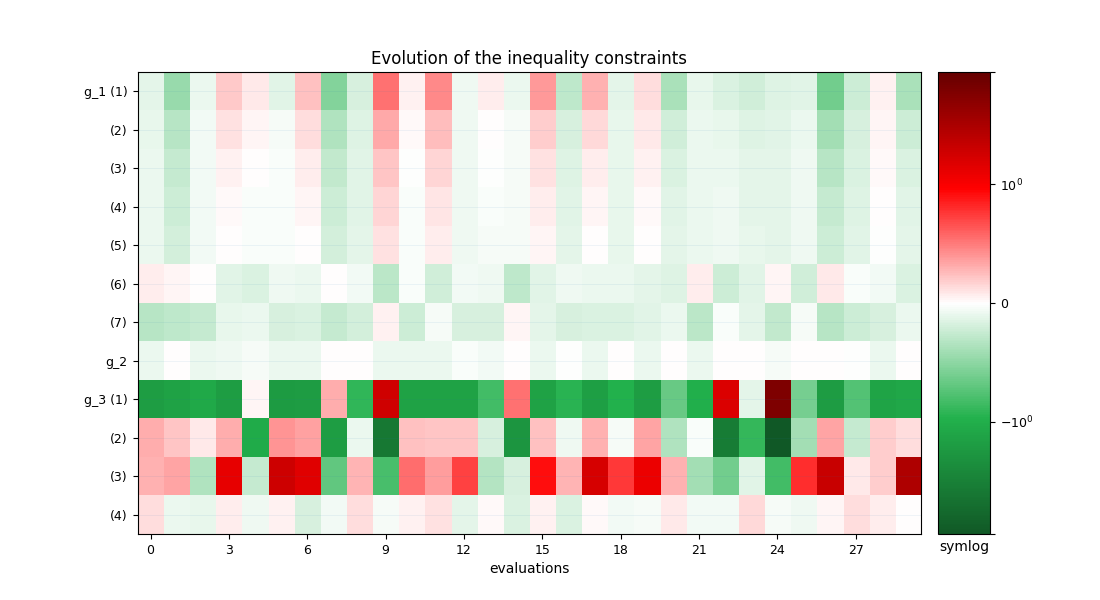
Plot the optimization history view¶
scenario.post_process("OptHistoryView", show=True, save=False)
Out:
/home/docs/checkouts/readthedocs.org/user_builds/gemseo/conda/3.0.3/lib/python3.8/site-packages/gemseo/post/opt_history_view.py:312: UserWarning: FixedFormatter should only be used together with FixedLocator
ax1.set_yticklabels(y_labels)
/home/docs/checkouts/readthedocs.org/user_builds/gemseo/conda/3.0.3/lib/python3.8/site-packages/gemseo/post/opt_history_view.py:716: MatplotlibDeprecationWarning: default base will change from np.e to 10 in 3.4. To suppress this warning specify the base keyword argument.
norm=SymLogNorm(linthresh=linthresh, vmin=-vmax, vmax=vmax),
/home/docs/checkouts/readthedocs.org/user_builds/gemseo/conda/3.0.3/lib/python3.8/site-packages/gemseo/post/opt_history_view.py:626: MatplotlibDeprecationWarning: default base will change from np.e to 10 in 3.4. To suppress this warning specify the base keyword argument.
norm=SymLogNorm(linthresh=1.0, vmin=-vmax, vmax=vmax),
/home/docs/checkouts/readthedocs.org/user_builds/gemseo/conda/3.0.3/lib/python3.8/site-packages/gemseo/post/opt_history_view.py:619: MatplotlibDeprecationWarning: Passing parameters norm and vmin/vmax simultaneously is deprecated since 3.3 and will become an error two minor releases later. Please pass vmin/vmax directly to the norm when creating it.
im1 = ax1.imshow(
<gemseo.post.opt_history_view.OptHistoryView object at 0x7fc298a2f670>
Plot the scatter matrix¶
scenario.post_process(
"ScatterPlotMatrix", show=True, save=False, variables_list=["y_4", "x_shared"]
)

Out:
<gemseo.post.scatter_mat.ScatterPlotMatrix object at 0x7fc299d70460>
Plot correlations¶
scenario.post_process("Correlations", show=True, save=False)
Out:
<gemseo.post.correlations.Correlations object at 0x7fc298a62670>
Total running time of the script: ( 0 minutes 3.925 seconds)