Note
Go to the end to download the full example code
Discipline¶
from __future__ import annotations
from numpy import array
from gemseo import configure_logger
from gemseo import create_discipline
from gemseo import generate_coupling_graph
from gemseo import generate_n2_plot
from gemseo import get_available_disciplines
from gemseo import get_discipline_inputs_schema
from gemseo import get_discipline_options_defaults
from gemseo import get_discipline_options_schema
from gemseo import get_discipline_outputs_schema
from gemseo.core.discipline import MDODiscipline
from gemseo.disciplines.utils import get_all_inputs
from gemseo.disciplines.utils import get_all_outputs
configure_logger()
<RootLogger root (INFO)>
In this example, we will discover the different functions of the API
related to disciplines, which are the GEMSEO’ objects
dedicated to the representation of an input-output process. All classes
implementing disciplines inherit from MDODiscipline which is an
abstract class.
Get available disciplines¶
The get_available_disciplines() function
can list the available disciplines:
get_available_disciplines()
['Aerodynamics', 'AnalyticDiscipline', 'AutoPyDiscipline', 'BaseDiscipline', 'CalibrationScenario', 'Calibrator', 'Concatenater', 'ConstraintAggregation', 'DOEScenario', 'DensityFilter', 'DiscFromExe', 'FilteringDiscipline', 'FininiteElementAnalysis', 'IshigamiDiscipline', 'JEPJavaDiscipline', 'JavaDiscipline', 'JobSchedulerDisciplineWrapper', 'LSF', 'LinearCombination', 'LinearDiscipline', 'MDOAdditiveChain', 'MDOChain', 'MDODiscipline', 'MDOObjectiveScenarioAdapter', 'MDOParallelChain', 'MDOScenario', 'MDOScenarioAdapter', 'MDOWarmStartedChain', 'MainDiscipline', 'MaterialModelInterpolation', 'MatlabDiscipline', 'Mission', 'PropaneComb1', 'PropaneComb2', 'PropaneComb3', 'PropaneReaction', 'RemappingDiscipline', 'RosenMF', 'SLURM', 'ScalableDiscipline', 'Scenario', 'ScilabDiscipline', 'Sellar1', 'Sellar2', 'SellarSystem', 'SobieskiAerodynamics', 'SobieskiAerodynamicsSG', 'SobieskiChain', 'SobieskiDiscipline', 'SobieskiDisciplineWithSimpleGrammar', 'SobieskiMDAGaussSeidel', 'SobieskiMDAJacobi', 'SobieskiMission', 'SobieskiMissionSG', 'SobieskiPropulsion', 'SobieskiPropulsionSG', 'SobieskiStructure', 'SobieskiStructureSG', 'Splitter', 'Structure', 'SurrogateDiscipline', 'TaylorDiscipline', 'VolumeFraction', 'XLSDiscipline']
Create a discipline¶
The create_discipline() function can create a
MDODiscipline or a list of MDODiscipline
by using its class name. Specific **options can be provided in
argument. E.g.
disciplines = create_discipline([
"SobieskiPropulsion",
"SobieskiAerodynamics",
"SobieskiMission",
"SobieskiStructure",
])
type(disciplines), type(disciplines[0]), isinstance(disciplines[0], MDODiscipline)
(<class 'list'>, <class 'gemseo.problems.mdo.sobieski.disciplines.SobieskiPropulsion'>, True)
This function can also be used to create a particular MDODiscipline
from scratch, such as AnalyticDiscipline
or AutoPyDiscipline. E.g.
addition = create_discipline("AnalyticDiscipline", expressions={"y": "x1+x2"})
addition.execute({"x1": array([1.0]), "x2": array([2.0])})
{'x1': array([1.]), 'x2': array([2.]), 'y': array([3.])}
Get all inputs/outputs¶
The get_all_inputs() function can list all the inputs
of a list of disciplines, including the sub-disciplines if the
argument recursive (default: False) is True,
merging the input data from the discipline grammars. E.g.
get_all_inputs(disciplines)
['c_0', 'c_1', 'c_2', 'c_3', 'c_4', 'x_1', 'x_2', 'x_3', 'x_shared', 'y_12', 'y_14', 'y_21', 'y_23', 'y_24', 'y_31', 'y_32', 'y_34']
The get_all_outputs() function can list all the inputs
of a list of disciplines, including the sub-disciplines if the
argument recursive (default: False) is True,
merging the input data from the discipline grammars. E.g.
get_all_outputs(disciplines)
['g_1', 'g_2', 'g_3', 'y_1', 'y_11', 'y_12', 'y_14', 'y_2', 'y_21', 'y_23', 'y_24', 'y_3', 'y_31', 'y_32', 'y_34', 'y_4']
Get discipline schemas for inputs, outputs and options¶
The function
get_discipline_inputs_schema()returns the inputs of a discipline. E.g.
get_discipline_inputs_schema(disciplines[0])
{'$schema': 'http://json-schema.org/draft-04/schema', 'type': 'object', 'properties': {'y_23': {'type': 'array', 'items': {'type': 'number'}}, 'x_3': {'type': 'array', 'items': {'type': 'number'}}, 'x_shared': {'type': 'array', 'items': {'type': 'number'}}, 'c_3': {'description': 'engine weight reference', 'type': 'array', 'items': {'type': 'number'}}}, 'required': ['c_3', 'x_3', 'x_shared', 'y_23']}
The function
get_discipline_outputs_schema()returns the outputs of a discipline. E.g.
get_discipline_outputs_schema(disciplines[0])
{'$schema': 'http://json-schema.org/draft-04/schema', 'type': 'object', 'properties': {'y_32': {'type': 'array', 'items': {'type': 'number'}}, 'y_31': {'type': 'array', 'items': {'type': 'number'}}, 'g_3': {'type': 'array', 'items': {'type': 'number'}}, 'y_3': {'type': 'array', 'items': {'type': 'number'}}, 'y_34': {'type': 'array', 'items': {'type': 'number'}}}, 'required': ['g_3', 'y_3', 'y_31', 'y_32', 'y_34']}
The function
get_discipline_options_schema()returns the options of a discipline. E.g.
get_discipline_options_schema("SobieskiMission")
{'$schema': 'http://json-schema.org/draft-04/schema', 'type': 'object', 'properties': {'linearization_mode': {'description': 'Linearization mode', 'enum': ['auto', 'direct', 'reverse', 'adjoint'], 'type': 'string'}, 'jac_approx_type': {'description': 'Jacobian approximation type', 'enum': ['finite_differences', 'complex_step'], 'type': 'string'}, 'jax_approx_step': {'minimum': 0, 'exclusiveMinimum': True, 'description': 'Step for finite differences or complex step for Jacobian approximation', 'type': 'number'}, 'jac_approx_use_threading': {'description': 'if True, use Threads instead of processes\n to parallelize the execution. \nMultiprocessing will serialize all the disciplines, \nwhile multithreading will share all the memory.\n This is important to note if you want to execute the same\n discipline multiple times, you shall use multiprocessing', 'type': 'boolean'}, 'jac_approx_wait_time': {'description': 'Time waited between two forks of the process or thread when using parallel jacobian approximations (parallel=True)', 'minimum': 0, 'type': 'number'}, 'jac_approx_n_processes': {'minimum': 1, 'description': 'maximum number of processors or threads on \nwhich the jacobian approximation is performed\n by default, 1 means no parallel calculations', 'type': 'integer'}, 'cache_type': {'description': 'Type of cache to be used. \nBy default, simple cache stores the last execution inputs and outputs \nin memory only to avoid computation of the outputs if the inputs are identical.\n To store more executions, use HDF5 caches, which stores data on the disk.\n There is a hashing mechanism which avoids reading on the disk for every calculation.', 'type': 'string'}, 'cache_tolerance': {'minimum': 0, 'description': 'Numerical tolerance on the relative norm of input vectors \n to consider that two sets of inputs are equal, and that the outputs may therefore be returned from the cache without calculations.', 'type': 'number'}, 'cache_hdf_file': {'format': 'uri', 'description': 'Path to the HDF5 file to store the cache data.', 'type': 'string'}, 'cache_hdf_node_path': {'description': 'Name of the HDF dataset to store the discipline\n data. If ``None``, the discipline name is used.', 'type': 'string'}, 'dtype': {'description': 'The data type for the NumPy arrays, either "float64" or "complex128".', 'type': 'string'}, 'enable_delay': {'description': 'If ``True``, wait one second before computation. If a positive number, wait the corresponding number of seconds. If ``False``, compute directly.', 'type': 'boolean'}}}
The function
get_discipline_options_defaults()can get the default option values of a discipline. E.g.
get_discipline_options_defaults("SobieskiMission")
{'dtype': <DataType.FLOAT: 'float64'>, 'enable_delay': False}
Plot coupling structure¶
The generate_coupling_graph() function plots the
coupling graph of a set of MDODiscipline:
generate_coupling_graph(disciplines, file_path="full_coupling_graph.pdf")
generate_coupling_graph(
disciplines, file_path="condensed_coupling_graph.pdf", full=False
)
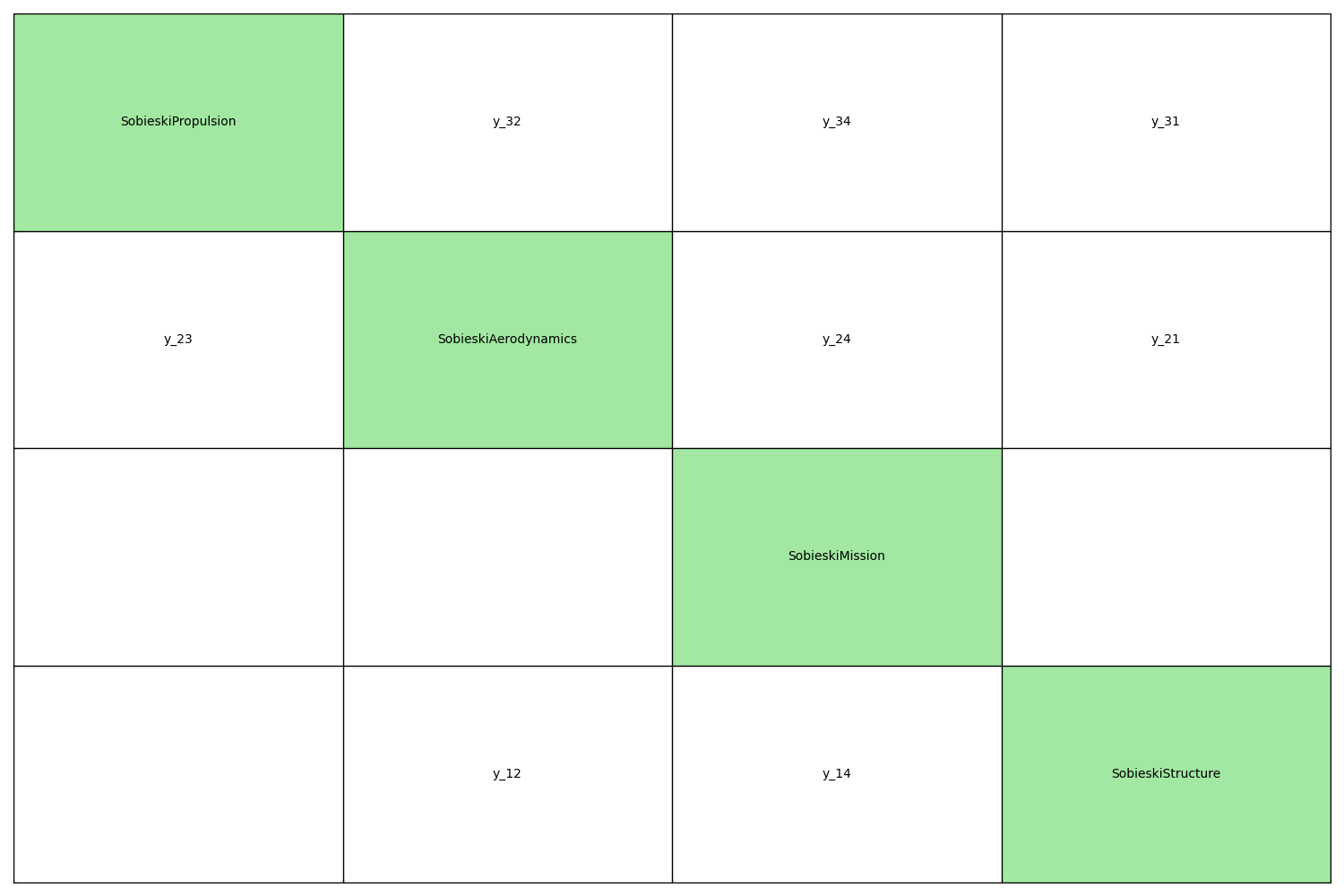
The generate_n2_plot() function plots the N2 diagram of
a set of MDODiscipline:
generate_n2_plot(disciplines, save=False, show=True)
Total running time of the script: (0 minutes 0.406 seconds)
