Note
Click here to download the full example code
Iris dataset¶
Presentation¶
This is one of the best known dataset to be found in the machine learning literature.
It was introduced by the statistician Ronald Fisher in his 1936 paper “The use of multiple measurements in taxonomic problems”, Annals of Eugenics. 7 (2): 179–188.
It contains 150 instances of iris plants:
50 Iris Setosa,
50 Iris Versicolour,
50 Iris Virginica.
Each instance is characterized by:
its sepal length in cm,
its sepal width in cm,
its petal length in cm,
its petal width in cm.
This dataset can be used for either clustering purposes or classification ones.
from __future__ import absolute_import, division, print_function, unicode_literals
from future import standard_library
from numpy.random import choice
from gemseo.api import configure_logger, load_dataset
configure_logger()
standard_library.install_aliases()
Load Iris dataset¶
We can easily load this dataset by means of the
load_dataset() function of the API:
iris = load_dataset("IrisDataset")
and get some information about it
print(iris)
Out:
Iris
| Number of samples: 150
| Number of variables: 5
| Variables names and sizes by group:
| - parameters: sepal_length (1), sepal_width (1), petal_length (1), petal_width (1)
| - labels: specy (1)
| Number of dimensions (total = 5) by group:
| - parameters: 4
| - labels: 1
Manipulate the dataset¶
We randomly select 10 samples to display.
shown_samples = choice(iris.length, size=10, replace=False)
If the pandas library is installed, we can export the iris dataset to a dataframe and print(it.
dataframe = iris.export_to_dataframe()
print(dataframe)
Out:
parameters labels
sepal_length sepal_width petal_length petal_width specy
0 0 0 0 0
0 5.1 3.5 1.4 0.2 0.0
1 4.9 3.0 1.4 0.2 0.0
2 4.7 3.2 1.3 0.2 0.0
3 4.6 3.1 1.5 0.2 0.0
4 5.0 3.6 1.4 0.2 0.0
.. ... ... ... ... ...
145 6.7 3.0 5.2 2.3 2.0
146 6.3 2.5 5.0 1.9 2.0
147 6.5 3.0 5.2 2.0 2.0
148 6.2 3.4 5.4 2.3 2.0
149 5.9 3.0 5.1 1.8 2.0
[150 rows x 5 columns]
We can also easily access the 10 samples previously selected, either globally
data = iris.get_all_data(False)
print(data[0][shown_samples, :])
Out:
[[5.8 4. 1.2 0.2 0. ]
[5.1 2.5 3. 1.1 1. ]
[6.6 3. 4.4 1.4 1. ]
[5.4 3.9 1.3 0.4 0. ]
[7.9 3.8 6.4 2. 2. ]
[6.3 3.3 4.7 1.6 1. ]
[6.9 3.1 5.1 2.3 2. ]
[5.1 3.8 1.9 0.4 0. ]
[4.7 3.2 1.6 0.2 0. ]
[6.9 3.2 5.7 2.3 2. ]]
or only the parameters:
parameters = iris.get_data_by_group("parameters")
print(parameters[shown_samples, :])
Out:
[[5.8 4. 1.2 0.2]
[5.1 2.5 3. 1.1]
[6.6 3. 4.4 1.4]
[5.4 3.9 1.3 0.4]
[7.9 3.8 6.4 2. ]
[6.3 3.3 4.7 1.6]
[6.9 3.1 5.1 2.3]
[5.1 3.8 1.9 0.4]
[4.7 3.2 1.6 0.2]
[6.9 3.2 5.7 2.3]]
or only the labels:
labels = iris.get_data_by_group("labels")
print(labels[shown_samples, :])
Out:
[[0.]
[1.]
[1.]
[0.]
[2.]
[1.]
[2.]
[0.]
[0.]
[2.]]
Plot the dataset¶
Lastly, we can plot the dataset in various ways. We will note that the samples are colored according to their labels.
Plot scatter matrix¶
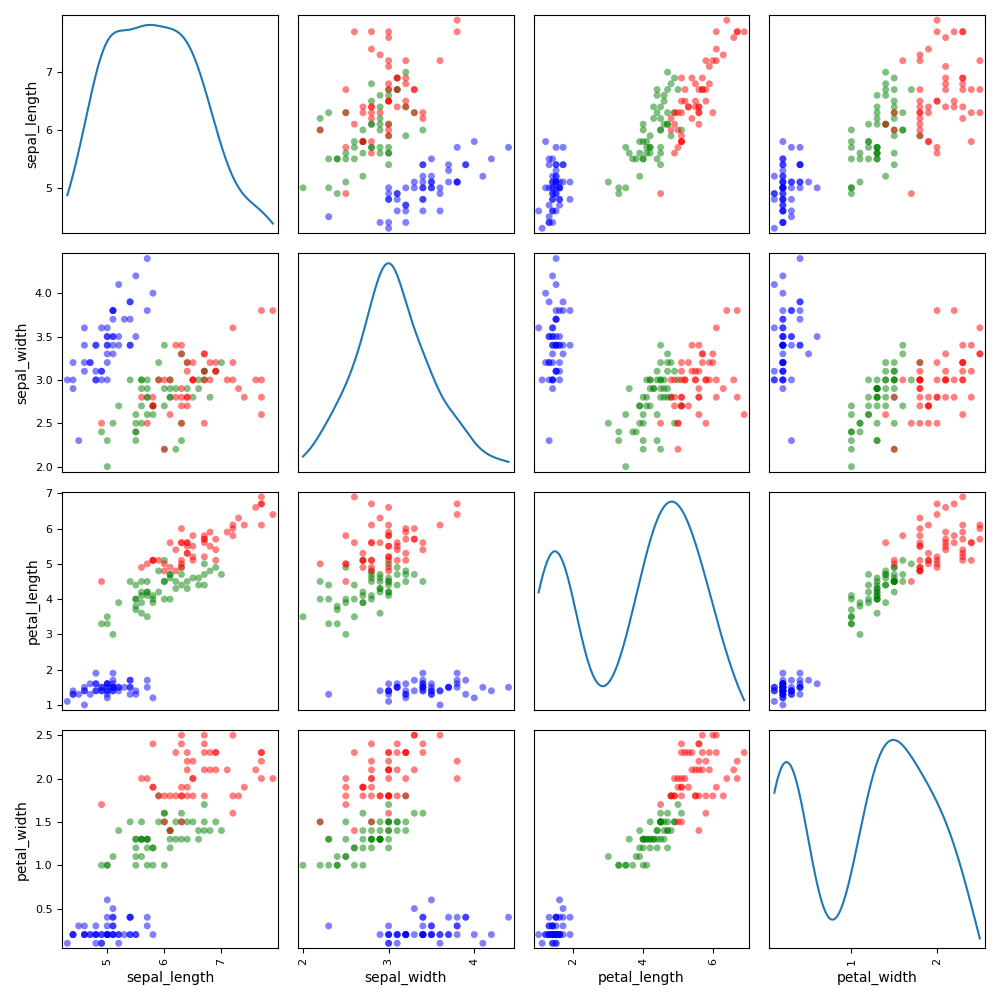
We can use the ScatterMatrix plot where each non-diagonal block
represents the samples according to the x- and y- coordinates names
while the diagonal ones approximate the probability distributions of the
variables, using either an histogram or a kernel-density estimator.
iris.plot("ScatterMatrix", classifier="specy", kde=True)
Out:
<gemseo.post.dataset.scatter_plot_matrix.ScatterMatrix object at 0x7fc29e2dc8e0>
Plot parallel coordinates¶
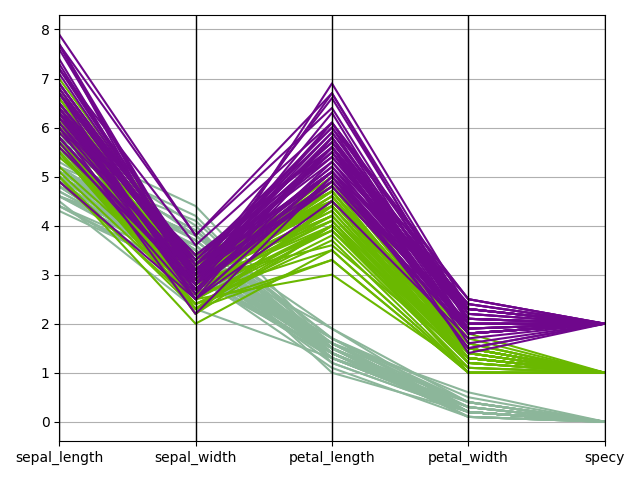
We can use the
ParallelCoordinates plot,
a.k.a. cowebplot, where each samples
is represented by a continuous straight line in pieces whose nodes are
indexed by the variables names and measure the variables values.
iris.plot("ParallelCoordinates", classifier="specy")
Out:
<gemseo.post.dataset.parallel_coordinates.ParallelCoordinates object at 0x7fc29de975b0>
Plot Andrews curves¶
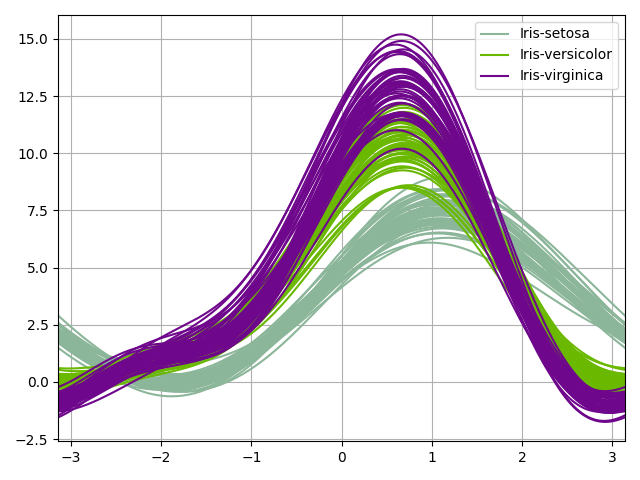
We can use the AndrewsCurves plot
which can be viewed as a smooth
version of the parallel coordinates. Each sample is represented by a curve
and if there is structure in data, it may be visible in the plot.
iris.plot("AndrewsCurves", classifier="specy")
Out:
<gemseo.post.dataset.andrews_curves.AndrewsCurves object at 0x7fc29e2dc790>