Note
Click here to download the full example code
Scalable diagonal discipline¶
Let us consider the
SobieskiAerodynamics discipline.
We want to build its ScalableDiscipline counterpart,
using a ScalableDiagonalModel
For that, we can use a 20-length DiagonalDOE
and test different sizes of variables or different settings
for the scalable diagonal discipline.
from __future__ import annotations
from gemseo.api import configure_logger
from gemseo.api import create_discipline
from gemseo.api import create_scalable
from gemseo.api import create_scenario
from gemseo.problems.sobieski.core.problem import SobieskiProblem
from matplotlib import pyplot as plt
Import¶
configure_logger()
<RootLogger root (INFO)>
Learning dataset¶
The first step is to build an AbstractFullCache dataset
from a DiagonalDOE.
Instantiate the discipline¶
For that, we instantiate the
SobieskiAerodynamics discipline
and set it up to cache all evaluations.
discipline = create_discipline("SobieskiAerodynamics")
Get the input space¶
We also define the input space on which to sample the discipline.
input_space = SobieskiProblem().design_space
input_names = [name for name in discipline.get_input_data_names() if name != "c_4"]
input_space.filter(input_names)
<gemseo.algos.design_space.DesignSpace object at 0x7f3c13708ac0>
Build the DOE scenario¶
Lastly, we sample the discipline by means of a DOEScenario
relying on both discipline and input space.
In order to build a diagonal scalable discipline,
a DiagonalDOE must be used.
scenario = create_scenario(
[discipline], "DisciplinaryOpt", "y_2", input_space, scenario_type="DOE"
)
for output_name in discipline.get_output_data_names():
if output_name != "y_2":
scenario.add_observable(output_name)
scenario.execute({"algo": "DiagonalDOE", "n_samples": 20})
INFO - 14:44:46:
INFO - 14:44:46: *** Start DOEScenario execution ***
INFO - 14:44:46: DOEScenario
INFO - 14:44:46: Disciplines: SobieskiAerodynamics
INFO - 14:44:46: MDO formulation: DisciplinaryOpt
INFO - 14:44:46: Optimization problem:
INFO - 14:44:46: minimize y_2(x_shared, x_2, y_32, y_12)
INFO - 14:44:46: with respect to x_2, x_shared, y_12, y_32
INFO - 14:44:46: over the design space:
INFO - 14:44:46: +-------------+-------------+--------------------+-------------+-------+
INFO - 14:44:46: | name | lower_bound | value | upper_bound | type |
INFO - 14:44:46: +-------------+-------------+--------------------+-------------+-------+
INFO - 14:44:46: | x_shared[0] | 0.01 | 0.05 | 0.09 | float |
INFO - 14:44:46: | x_shared[1] | 30000 | 45000 | 60000 | float |
INFO - 14:44:46: | x_shared[2] | 1.4 | 1.6 | 1.8 | float |
INFO - 14:44:46: | x_shared[3] | 2.5 | 5.5 | 8.5 | float |
INFO - 14:44:46: | x_shared[4] | 40 | 55 | 70 | float |
INFO - 14:44:46: | x_shared[5] | 500 | 1000 | 1500 | float |
INFO - 14:44:46: | x_2 | 0.75 | 1 | 1.25 | float |
INFO - 14:44:46: | y_32 | 0.235 | 0.5027962499999999 | 0.795 | float |
INFO - 14:44:46: | y_12[0] | 24850 | 50606.9742 | 77250 | float |
INFO - 14:44:46: | y_12[1] | 0.45 | 0.95 | 1.5 | float |
INFO - 14:44:46: +-------------+-------------+--------------------+-------------+-------+
INFO - 14:44:46: Solving optimization problem with algorithm DiagonalDOE:
INFO - 14:44:46: ... 0%| | 0/20 [00:00<?, ?it]
INFO - 14:44:46: ... 100%|██████████| 20/20 [00:00<00:00, 451.45 it/sec, obj=[7.72500000e+04 1.27392070e+04 6.06395673e+00]]
INFO - 14:44:46: Optimization result:
INFO - 14:44:46: Optimizer info:
INFO - 14:44:46: Status: None
INFO - 14:44:46: Message: None
INFO - 14:44:46: Number of calls to the objective function by the optimizer: 20
INFO - 14:44:46: Solution:
INFO - 14:44:46: Objective: 25508.372961119574
INFO - 14:44:46: Design space:
INFO - 14:44:46: +-------------+-------------+-------+-------------+-------+
INFO - 14:44:46: | name | lower_bound | value | upper_bound | type |
INFO - 14:44:46: +-------------+-------------+-------+-------------+-------+
INFO - 14:44:46: | x_shared[0] | 0.01 | 0.01 | 0.09 | float |
INFO - 14:44:46: | x_shared[1] | 30000 | 30000 | 60000 | float |
INFO - 14:44:46: | x_shared[2] | 1.4 | 1.4 | 1.8 | float |
INFO - 14:44:46: | x_shared[3] | 2.5 | 2.5 | 8.5 | float |
INFO - 14:44:46: | x_shared[4] | 40 | 40 | 70 | float |
INFO - 14:44:46: | x_shared[5] | 500 | 500 | 1500 | float |
INFO - 14:44:46: | x_2 | 0.75 | 0.75 | 1.25 | float |
INFO - 14:44:46: | y_32 | 0.235 | 0.235 | 0.795 | float |
INFO - 14:44:46: | y_12[0] | 24850 | 24850 | 77250 | float |
INFO - 14:44:46: | y_12[1] | 0.45 | 0.45 | 1.5 | float |
INFO - 14:44:46: +-------------+-------------+-------+-------------+-------+
INFO - 14:44:46: *** End DOEScenario execution (time: 0:00:00.059042) ***
{'eval_jac': False, 'algo': 'DiagonalDOE', 'n_samples': 20}
Scalable diagonal discipline¶
Build the scalable discipline¶
The second step is to build a ScalableDiscipline,
using a ScalableDiagonalModel and the database
converted to a Dataset.
dataset = scenario.export_to_dataset(opt_naming=False)
scalable = create_scalable("ScalableDiagonalModel", dataset)
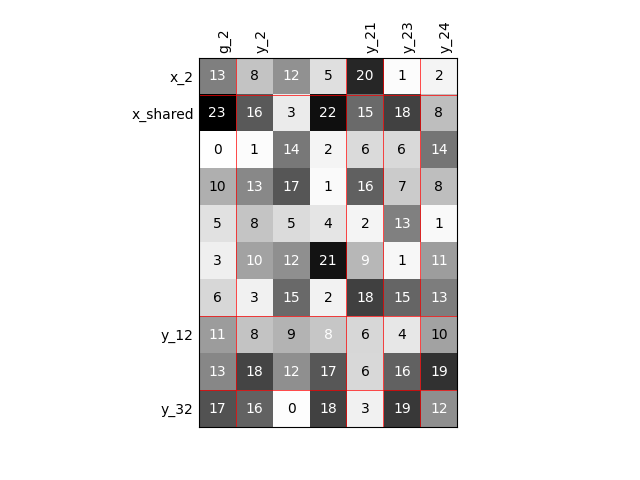
Visualize the input-output dependencies¶
We can easily access the underlying ScalableDiagonalModel
and plot the corresponding input-output dependency matrix
where the level of gray and the number (in [0,100]) represent
the degree of dependency between inputs and outputs.
Input are on the left while outputs are at the top.
More precisely, for a given output component located at the top of the graph,
these degrees are contributions to the output component and they add up to 1.
In other words, a degree expresses this contribution in percentage
and for a given column, the elements add up to 100.
scalable.scalable_model.plot_dependency(save=False, show=False)
'None'
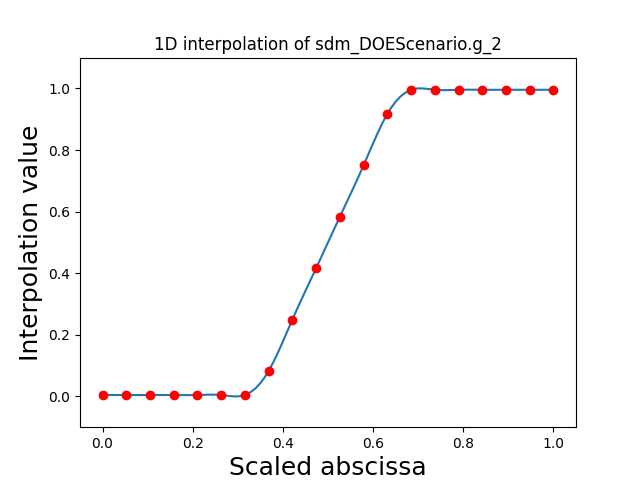
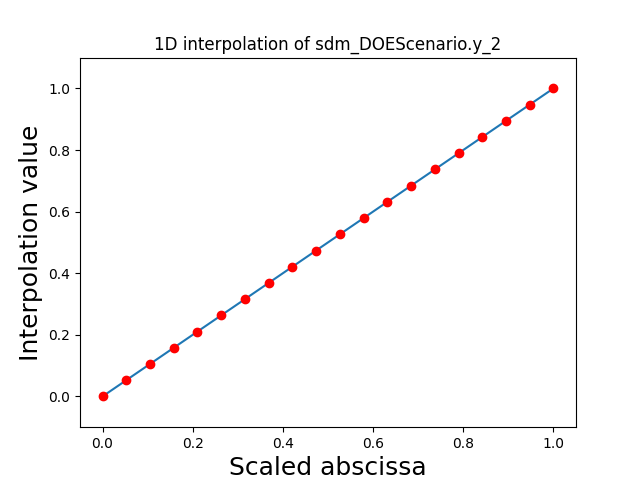
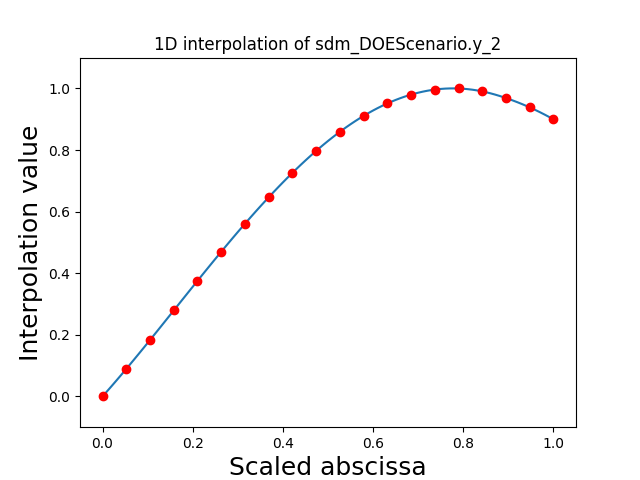
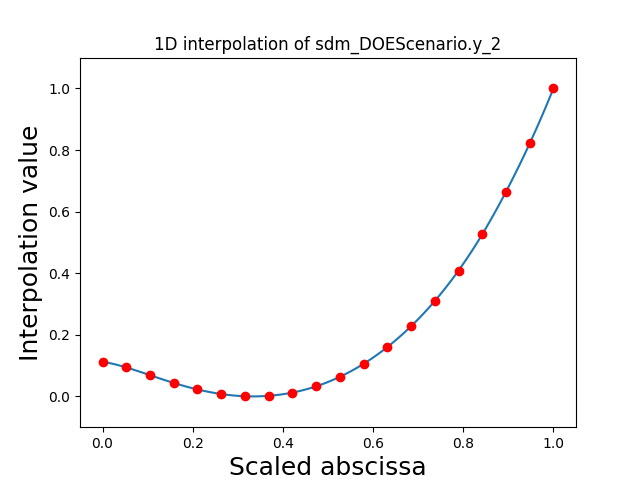
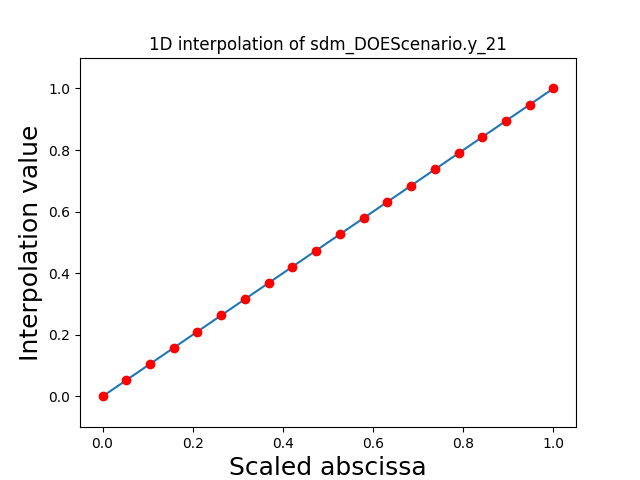
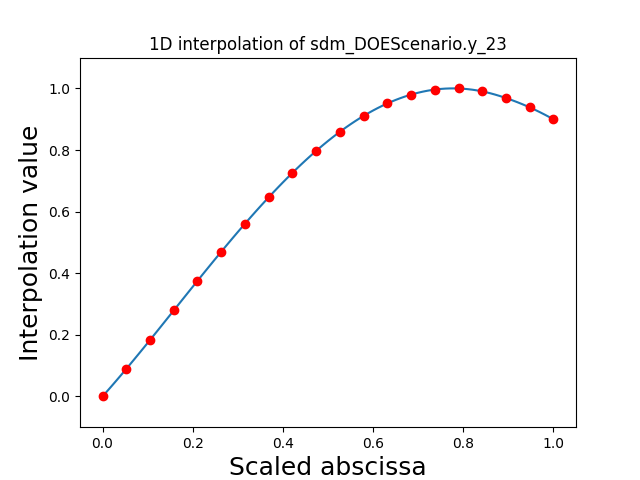
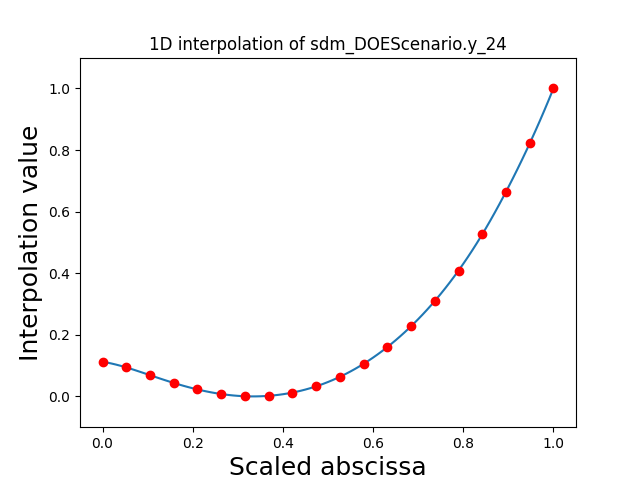
Visualize the 1D interpolations¶
For every output, we can also visualize a spline interpolation of the output samples over the diagonal of the input space.
scalable.scalable_model.plot_1d_interpolations(save=False, show=False)
[]
Increased problem dimension¶
We can repeat the construction of the scalable discipline for different sizes of variables and visualize the input-output dependency matrices.
Twice as many inputs¶
For example, we can increase the size of each input by a factor of 2.
sizes = {name: dataset.sizes[name] * 2 for name in input_names}
scalable = create_scalable("ScalableDiagonalModel", dataset, sizes)
scalable.scalable_model.plot_dependency(save=False, show=False)
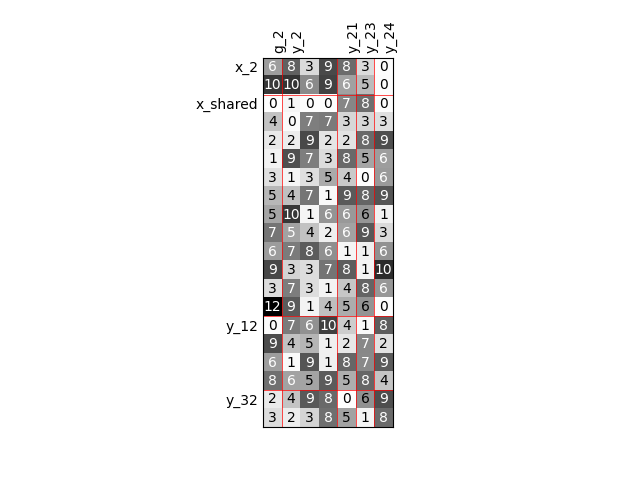
'None'
Twice as many outputs¶
Or we can increase the size of each output by a factor of 2.
sizes = {
name: discipline.cache.names_to_sizes[name] * 2
for name in discipline.get_output_data_names()
}
scalable = create_scalable("ScalableDiagonalModel", dataset, sizes)
scalable.scalable_model.plot_dependency(save=False, show=False)
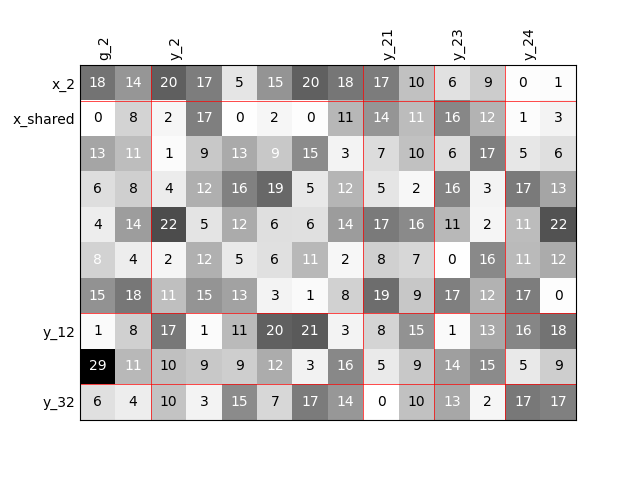
'None'
Twice as many variables¶
Or we can increase the size of each input and each output by a factor of 2.
names = input_names + list(discipline.get_output_data_names())
sizes = {name: dataset.sizes[name] * 2 for name in names}
scalable = create_scalable("ScalableDiagonalModel", dataset, sizes)
scalable.scalable_model.plot_dependency(save=False, show=False)
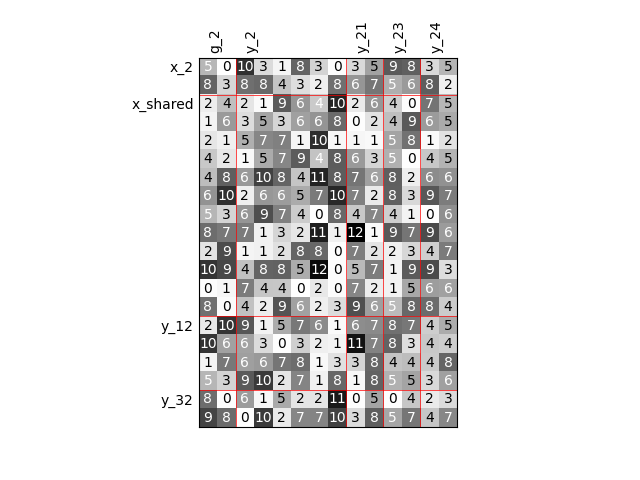
'None'
Binary IO dependencies¶
By default, any output component depends on any input component with a random level. We can also consider sparser input-output dependency by means of binary input-output dependency matrices. For that, we have to set the value of the fill factor which represents the part of connection between inputs and outputs. Then, a connection is represented by a black square while an absence of connection is presented by a white one. When the fill factor is equal to 1, any input is connected to any output. Conversely, when the fill factor is equal to 0, there is not a single connection between inputs and outputs.
Fill factor = 0.2¶
scalable = create_scalable("ScalableDiagonalModel", dataset, sizes, fill_factor=0.2)
scalable.scalable_model.plot_dependency(save=False, show=False)
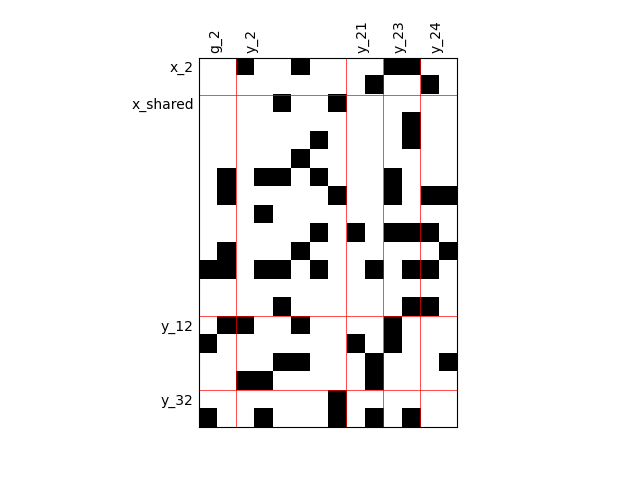
'None'
Fill factor = 0.5¶
scalable = create_scalable("ScalableDiagonalModel", dataset, sizes, fill_factor=0.5)
scalable.scalable_model.plot_dependency(save=False, show=False)
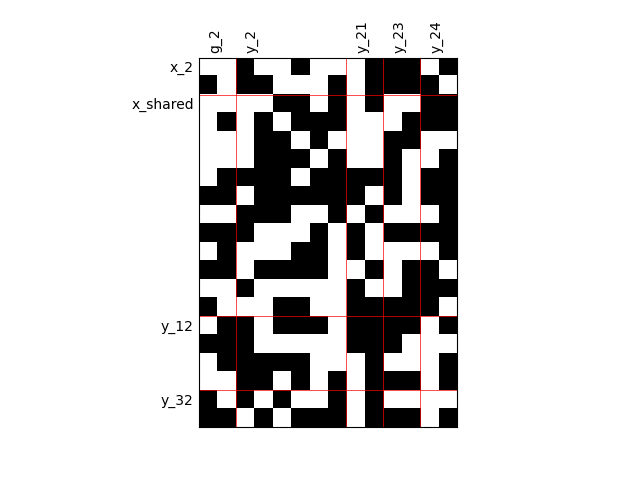
'None'
Fill factor = 0.8¶
scalable = create_scalable("ScalableDiagonalModel", dataset, sizes, fill_factor=0.8)
scalable.scalable_model.plot_dependency(save=False, show=False)
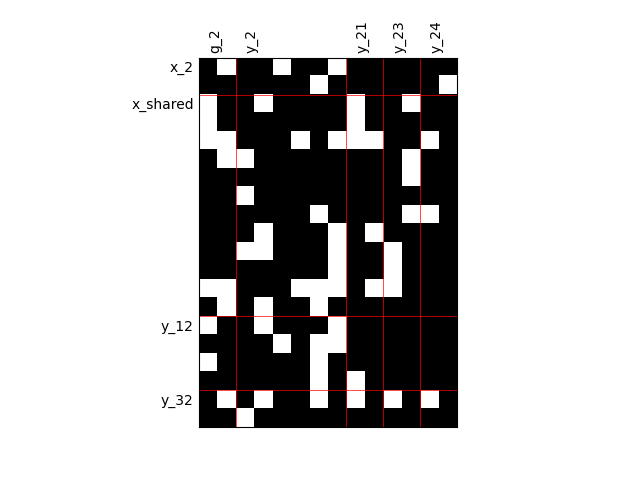
'None'
Heterogeneous dependencies¶
scalable = create_scalable(
"ScalableDiagonalModel", dataset, sizes, fill_factor={"y_2": 0.2}
)
scalable.scalable_model.plot_dependency(save=False, show=False)
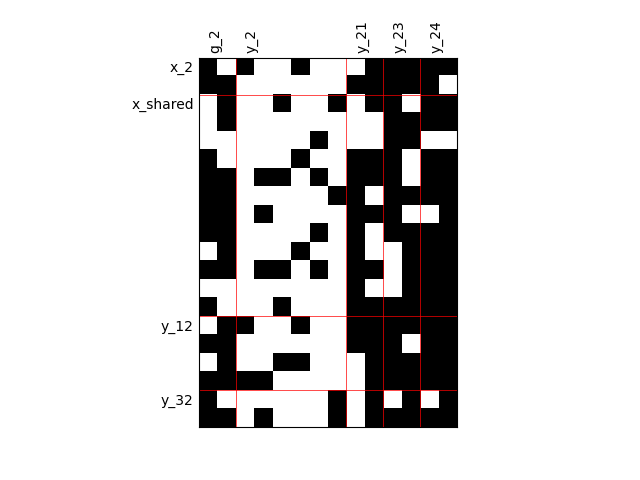
'None'
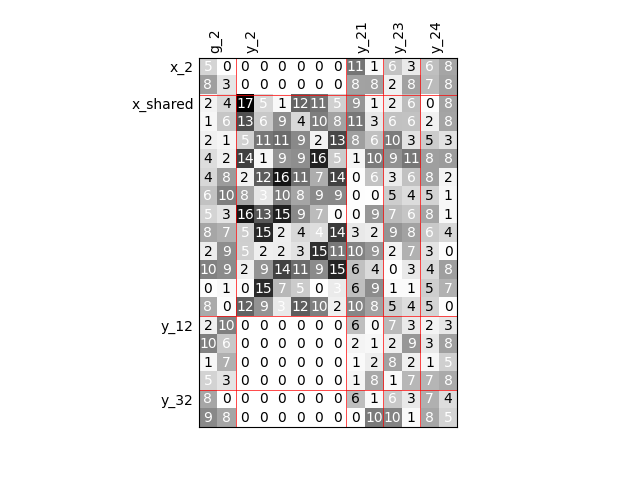
Group dependencies¶
scalable = create_scalable(
"ScalableDiagonalModel", dataset, sizes, group_dep={"y_2": ["x_shared"]}
)
scalable.scalable_model.plot_dependency(save=False, show=False)
# Workaround for HTML rendering, instead of ``show=True``
plt.show()

Total running time of the script: ( 0 minutes 3.288 seconds)