analysis module¶
Class for the estimation of various correlation coefficients.
- class gemseo.uncertainty.sensitivity.correlation.analysis.CorrelationAnalysis(disciplines, parameter_space, n_samples, output_names=None, algo=None, algo_options=None, formulation='MDF', **formulation_options)[source]
Bases:
SensitivityAnalysisSensitivity analysis based on indices using correlation measures.
Examples
>>> from numpy import pi >>> from gemseo import create_discipline, create_parameter_space >>> from gemseo.uncertainty.sensitivity.correlation.analysis import ( ... CorrelationAnalysis ... ) >>> >>> expressions = {"y": "sin(x1)+7*sin(x2)**2+0.1*x3**4*sin(x1)"} >>> discipline = create_discipline( ... "AnalyticDiscipline", expressions=expressions ... ) >>> >>> parameter_space = create_parameter_space() >>> parameter_space.add_random_variable( ... "x1", "OTUniformDistribution", minimum=-pi, maximum=pi ... ) >>> parameter_space.add_random_variable( ... "x2", "OTUniformDistribution", minimum=-pi, maximum=pi ... ) >>> parameter_space.add_random_variable( ... "x3", "OTUniformDistribution", minimum=-pi, maximum=pi ... ) >>> >>> analysis = CorrelationAnalysis([discipline], parameter_space, n_samples=1000) >>> indices = analysis.compute_indices()
- Parameters:
disciplines (Collection[MDODiscipline]) – The discipline or disciplines to use for the analysis.
parameter_space (ParameterSpace) – A parameter space.
n_samples (int) – A number of samples. If
None, the number of samples is computed by the algorithm.output_names (Iterable[str] | None) – The disciplines’ outputs to be considered for the analysis. If
None, use all the outputs.algo (str | None) – The name of the DOE algorithm. If
None, use theSensitivityAnalysis.DEFAULT_DRIVER.algo_options (Mapping[str, DOELibraryOptionType] | None) – The options of the DOE algorithm.
formulation (str) –
The name of the
MDOFormulationto sample the disciplines.By default it is set to “MDF”.
**formulation_options (Any) – The options of the
MDOFormulation.
- class Method(value)[source]
Bases:
StrEnumThe names of the sensitivity methods.
- KENDALL = 'Kendall'
The Kendall rank correlation coefficient.
- PCC = 'PCC'
The partial correlation coefficient.
- PEARSON = 'Pearson'
The Pearson coefficient.
- PRCC = 'PRCC'
The partial rank correlation coefficient.
- SPEARMAN = 'Spearman'
The Spearman coefficient.
- SRC = 'SRC'
The standard regression coefficient.
- SRRC = 'SRRC'
The standard rank regression coefficient.
- SSRC = 'SSRC'
The squared standard regression coefficient.
- compute_indices(outputs=None)[source]
Compute the sensitivity indices.
- Parameters:
outputs (str | Sequence[str] | None) – The output(s) for which to display the sensitivity indices. If
None, use the default outputs set at instantiation.- Returns:
The sensitivity indices.
With the following structure:
{ "method_name": { "output_name": [ { "input_name": data_array, } ] } }
- Return type:
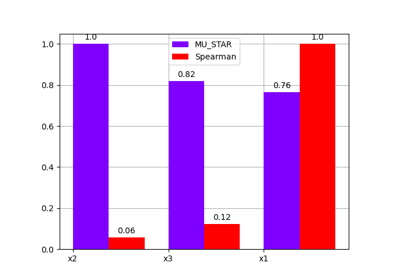
- plot(output, inputs=None, title=None, save=True, show=False, file_path=None, directory_path=None, file_name=None, file_format=None)[source]
Plot the sensitivity indices.
- Parameters:
output (VariableType) – The output for which to display sensitivity indices, either a name or a tuple of the form (name, component). If name, its first component is considered.
inputs (Iterable[str] | None) – The inputs to display. If None, display all.
title (str | None) – The title of the plot. If None, no title.
save (bool) –
If True, save the figure.
By default it is set to True.
show (bool) –
If True, show the figure.
By default it is set to False.
file_path (str | Path | None) – A file path. Either a complete file path, a directory name or a file name. If None, use a default file name and a default directory. The file extension is inferred from filepath extension, if any.
file_format (str | None) – A file format, e.g. ‘png’, ‘pdf’, ‘svg’, … Used when
file_pathdoes not have any extension. If None, use a default file extension.directory_path (str | Path | None) –
file_name (str | None) –
- Return type:
None
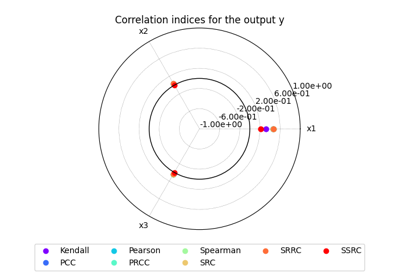
- plot_radar(outputs, inputs=None, title=None, save=True, show=False, file_path=None, directory_path=None, file_name=None, file_format=None, min_radius=-1.0, max_radius=1.0, **options)[source]
Plot the sensitivity indices on a radar chart.
This method may consider one or more outputs, as well as all inputs (default behavior) or a subset.
For visualization purposes, it is also possible to change the minimum and maximum radius values.
- Parameters:
outputs (OutputsType) – The outputs for which to display sensitivity indices, either a name, a list of names, a (name, component) tuple, a list of such tuples or a list mixing such tuples and names. When a name is specified, all its components are considered. If None, use the default outputs.
inputs (Iterable[str] | None) – The inputs to display. If None, display all.
standardize – If True, standardize the indices between 0 and 1 for each output.
title (str | None) – The title of the plot. If None, no title.
save (bool) –
If True, save the figure.
By default it is set to True.
show (bool) –
If True, show the figure.
By default it is set to False.
file_path (str | Path | None) – The path of the file to save the figures. If the extension is missing, use
file_extension. If None, create a file path fromdirectory_path,file_nameandfile_extension.directory_path (str | Path | None) – The path of the directory to save the figures. If None, use the current working directory.
file_name (str | None) – The name of the file to save the figures. If None, use a default one generated by the post-processing.
file_format (str | None) – A file extension, e.g. ‘png’, ‘pdf’, ‘svg’, … If None, use a default file extension.
min_radius (float) –
The minimal radial value. If None, from data.
By default it is set to -1.0.
max_radius (float) –
The maximal radial value. If None, from data.
By default it is set to 1.0.
options (bool) –
- Returns:
A radar chart representing the sensitivity indices.
- Return type:
- DEFAULT_DRIVER = 'OT_MONTE_CARLO'
- dataset: IODataset
The dataset containing the discipline evaluations.
- property indices: dict[str, Dict[str, List[Dict[str, numpy.ndarray]]]]
The sensitivity indices.
With the following structure:
{ "method_name": { "output_name": [ { "input_name": data_array, } ] } }
- property kendall: Dict[str, List[Dict[str, ndarray]]]
The Kendall rank correlation coefficients.
With the following structure:
{ "output_name": [ { "input_name": data_array, } ] }
- property pcc: Dict[str, List[Dict[str, ndarray]]]
The Partial Correlation Coefficients.
With the following structure:
{ "output_name": [ { "input_name": data_array, } ] }
- property pearson: Dict[str, List[Dict[str, ndarray]]]
The Pearson coefficients.
With the following structure:
{ "output_name": [ { "input_name": data_array, } ] }
- property prcc: Dict[str, List[Dict[str, ndarray]]]
The Partial Rank Correlation Coefficients.
With the following structure:
{ "output_name": [ { "input_name": data_array, } ] }
- property spearman: Dict[str, List[Dict[str, ndarray]]]
The Spearman coefficients.
ith the following structure:
{ "output_name": [ { "input_name": data_array, } ] }
- property src: Dict[str, List[Dict[str, ndarray]]]
The Standard Regression Coefficients.
With the following structure:
{ "output_name": [ { "input_name": data_array, } ] }