Note
Go to the end to download the full example code.
MSE for regression models#
from __future__ import annotations
from matplotlib import pyplot as plt
from numpy import array
from numpy import linspace
from numpy import newaxis
from numpy import sin
from gemseo.datasets.io_dataset import IODataset
from gemseo.mlearning.regression.algos.polyreg import PolynomialRegressor
from gemseo.mlearning.regression.algos.rbf import RBFRegressor
from gemseo.mlearning.regression.quality.mse_measure import MSEMeasure
Given a dataset \((x_i,y_i,\hat{y}_i)_{1\leq i \leq N}\) where \(x_i\) is an input point, \(y_i\) is an output observation and \(\hat{y}_i=\hat{f}(x_i)\) is an output prediction computed by a regression model \(\hat{f}\), the mean squared error (MSE) metric is written
The lower, the better.
From a quantitative point of view,
this depends on the order of magnitude of the outputs.
The square root of this average is often easier to interpret,
as it is expressed in the units of the output (see RMSEMeasure).
To illustrate this quality measure, let us consider the function \(f(x)=(6x-2)^2\sin(12x-4)\) [FSK08]:
def f(x):
return (6 * x - 2) ** 2 * sin(12 * x - 4)
and try to approximate it with a polynomial of order 3.
For this, we can take these 7 learning input points
x_train = array([0.1, 0.3, 0.5, 0.6, 0.8, 0.9, 0.95])
and evaluate the model f over this design of experiments (DOE):
y_train = f(x_train)
Then,
we create an IODataset from these 7 learning samples:
dataset_train = IODataset()
dataset_train.add_input_group(x_train[:, newaxis], ["x"])
dataset_train.add_output_group(y_train[:, newaxis], ["y"])
and build a PolynomialRegressor with degree=3 from it:
polynomial = PolynomialRegressor(dataset_train, degree=3)
polynomial.learn()
Before using it, we are going to measure its quality with the MSE metric:
mse = MSEMeasure(polynomial)
result = mse.compute_learning_measure()
result, result**0.5 / (y_train.max() - y_train.min())
(array([5.6443418]), array([0.137707]))
This result is medium (14% of the learning output range), and we can be expected to a poor generalization quality. As the cost of this academic function is zero, we can approximate this generalization quality with a large test dataset whereas the usual test size is about 20% of the training size.
x_test = linspace(0.0, 1.0, 100)
y_test = f(x_test)
dataset_test = IODataset()
dataset_test.add_input_group(x_test[:, newaxis], ["x"])
dataset_test.add_output_group(y_test[:, newaxis], ["y"])
result = mse.compute_test_measure(dataset_test)
result, result**0.5 / (y_test.max() - y_test.min())
(array([11.00451361]), array([0.15181886]))
The quality is higher than 15% of the test output range, which is pretty mediocre. This can be explained by a broader generalization domain than that of learning, which highlights the difficulties of extrapolation:
plt.plot(x_test, y_test, "-b", label="Reference")
plt.plot(x_train, y_train, "ob")
plt.plot(x_test, polynomial.predict(x_test[:, newaxis]), "-r", label="Prediction")
plt.plot(x_train, polynomial.predict(x_train[:, newaxis]), "or")
plt.legend()
plt.grid()
plt.show()
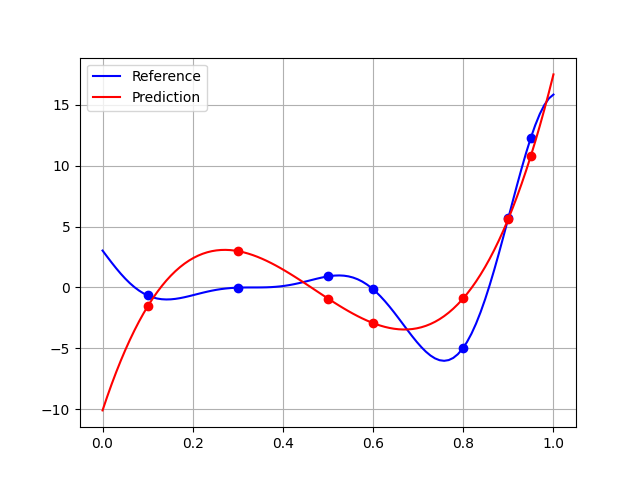
Using the learning domain would slightly improve the quality:
x_test = linspace(x_train.min(), x_train.max(), 100)
y_test_in_large_domain = y_test
y_test = f(x_test)
dataset_test_in_learning_domain = IODataset()
dataset_test_in_learning_domain.add_input_group(x_test[:, newaxis], ["x"])
dataset_test_in_learning_domain.add_output_group(y_test[:, newaxis], ["y"])
mse.compute_test_measure(dataset_test_in_learning_domain)
result, result**0.5 / (y_test.max() - y_test.min())
(array([11.00451361]), array([0.18111514]))
Lastly, to get better results without new learning points, we would have to change the regression model:
rbf = RBFRegressor(dataset_train)
rbf.learn()
The quality of this RBFRegressor is quite good,
both on the learning side:
mse_rbf = MSEMeasure(rbf)
result = mse_rbf.compute_learning_measure()
result, result**0.5 / (y_train.max() - y_train.min())
(array([7.61466172e-29]), array([5.05795166e-16]))
and on the validation side:
result = mse_rbf.compute_test_measure(dataset_test_in_learning_domain)
result, result**0.5 / (y_test.max() - y_test.min())
(array([0.02227183]), array([0.00814793]))
including the larger domain:
result = mse_rbf.compute_test_measure(dataset_test)
result, result**0.5 / (y_test_in_large_domain.max() - y_test_in_large_domain.min())
(array([0.29357082]), array([0.02479686]))
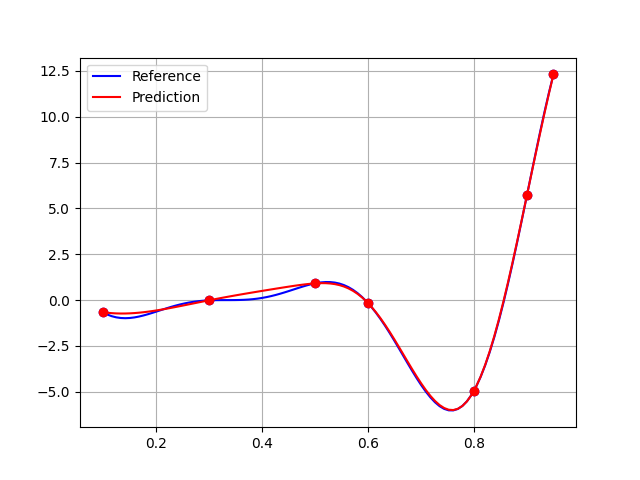
A final plot to convince us:
plt.plot(x_test, y_test, "-b", label="Reference")
plt.plot(x_train, y_train, "ob")
plt.plot(x_test, rbf.predict(x_test[:, newaxis]), "-r", label="Prediction")
plt.plot(x_train, rbf.predict(x_train[:, newaxis]), "or")
plt.legend()
plt.grid()
plt.show()
Total running time of the script: (0 minutes 0.112 seconds)
