Note
Go to the end to download the full example code.
Mixture of experts#
A MOERegressor is a mixture of experts for regression purposes.
In this demo, we load a dataset (the Rosenbrock function in 2D) and apply a mixture of experts regression model to obtain an approximation.
from __future__ import annotations
import matplotlib.pyplot as plt
from numpy import array
from numpy import hstack
from numpy import linspace
from numpy import meshgrid
from numpy import nonzero
from numpy import sqrt
from numpy import zeros
from gemseo import create_benchmark_dataset
from gemseo.mlearning import create_regression_model
Dataset (Rosenbrock)#
We here consider the Rosenbrock function with two inputs, on the interval \([-2, 2] \times [-2, 2]\).
Load dataset#
A prebuilt dataset for the Rosenbrock function with two inputs is given as a dataset parametrization, based on a full factorial DOE of the input space with 100 points.
dataset = create_benchmark_dataset("RosenbrockDataset", opt_naming=False)
Print information#
Information about the dataset can easily be displayed by printing the dataset directly.
dataset
Show dataset#
The dataset object can present the data in tabular form.
dataset
Mixture of experts (MoE)#
In this section we load a mixture of experts regression model through the machine learning API, using clustering, classification and regression models.
Mixture of experts model#
We construct the MoE model using the predefined parameters,
and fit the model to the dataset through the learn() method.
model = create_regression_model("MOERegressor", dataset)
model.set_clusterer("KMeans", n_clusters=3)
model.set_classifier("KNNClassifier", n_neighbors=5)
model.set_regressor("GaussianProcessRegressor")
model.learn()
Tests#
Here, we test the mixture of experts method applied to two points: (1, 1), the global minimum, where the function is zero, and (-2, -2), an extreme point where the function has a high value (max on the domain). The classes are expected to be different at the two points.
input_value = {"x": array([1, 1])}
another_input_value = {"x": array([[1, 1], [-2, -2]])}
for value in [input_value, another_input_value]:
print("Input value:", value)
print("Class:", model.predict_class(value))
print("Prediction:", model.predict(value))
print("Local model predictions:")
for cls in range(model.n_clusters):
print(f"Local model {cls}: {model.predict_local_model(value, cls)}")
print()
Input value: {'x': array([1, 1])}
Class: {'labels': array([0])}
Prediction: {'rosen': array([9.10887101])}
Local model predictions:
Local model 0: {'rosen': array([9.10887101])}
Local model 1: {'rosen': array([-40.29069491])}
Local model 2: {'rosen': array([315.46123093])}
Input value: {'x': array([[ 1, 1],
[-2, -2]])}
Class: {'labels': array([[0],
[2]])}
Prediction: {'rosen': array([[ 9.10887101],
[3608.99998212]])}
Local model predictions:
Local model 0: {'rosen': array([[ 9.10887101],
[553.30101764]])}
Local model 1: {'rosen': array([[ -40.29069491],
[1191.17575979]])}
Local model 2: {'rosen': array([[ 315.46123093],
[3608.99998212]])}
Plot clusters#
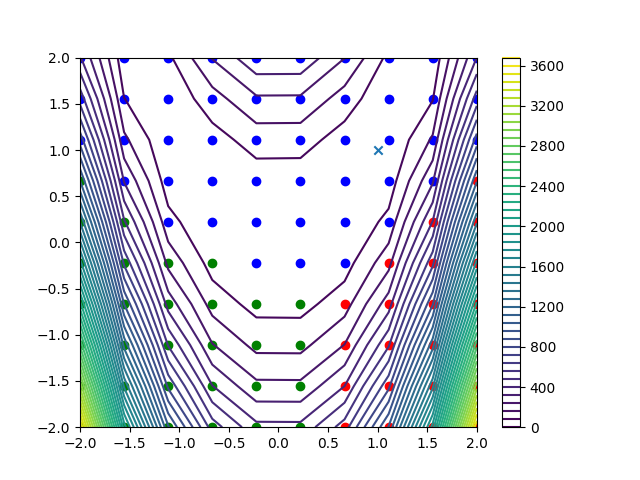
Here, we plot the 10x10 = 100 Rosenbrock function data points, with colors representing the obtained clusters. The Rosenbrock function is represented by a contour plot in the background.
n_samples = dataset.n_samples
# Dataset is based on a DOE of 100=10^2 fullfact.
input_dim = int(sqrt(n_samples))
assert input_dim**2 == n_samples # Check that n_samples is a square number
colors = ["b", "r", "g", "o", "y"]
inputs = dataset.input_dataset.to_numpy()
outputs = dataset.output_dataset.to_numpy()
x = inputs[:input_dim, 0]
y = inputs[:input_dim, 0]
Z = zeros((input_dim, input_dim))
for i in range(input_dim):
Z[i, :] = outputs[input_dim * i : input_dim * (i + 1), 0]
fig = plt.figure()
cnt = plt.contour(x, y, Z, 50)
fig.colorbar(cnt)
for index in range(model.n_clusters):
samples = nonzero(model.labels == index)[0]
plt.scatter(inputs[samples, 0], inputs[samples, 1], color=colors[index])
plt.scatter(1, 1, marker="x")
plt.show()
Plot data and predictions from final model#
We construct a refined input space, and compute the model predictions.
refinement = 200
fine_x = linspace(x[0], x[-1], refinement)
fine_y = linspace(y[0], y[-1], refinement)
fine_x, fine_y = meshgrid(fine_x, fine_y)
fine_input = {"x": hstack([fine_x.flatten()[:, None], fine_y.flatten()[:, None]])}
fine_z = model.predict(fine_input)
# Reshape
fine_z = fine_z["rosen"].reshape((refinement, refinement))
plt.figure()
plt.imshow(Z)
plt.colorbar()
plt.title("Original data")
plt.show()
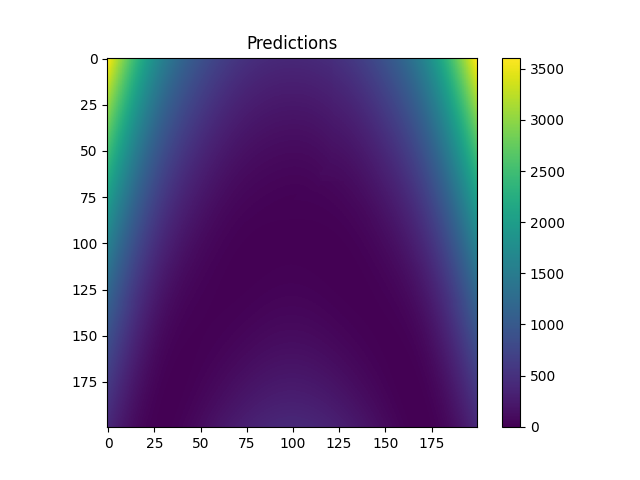
plt.figure()
plt.imshow(fine_z)
plt.colorbar()
plt.title("Predictions")
plt.show()
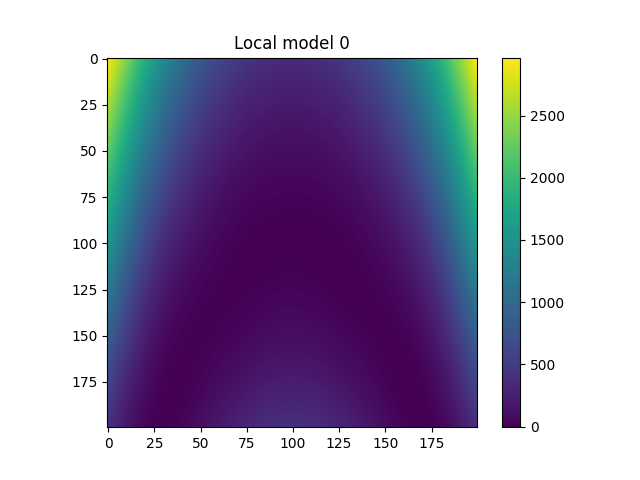
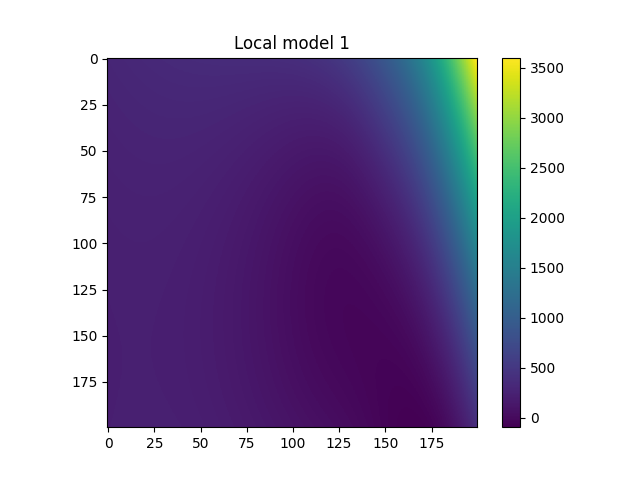
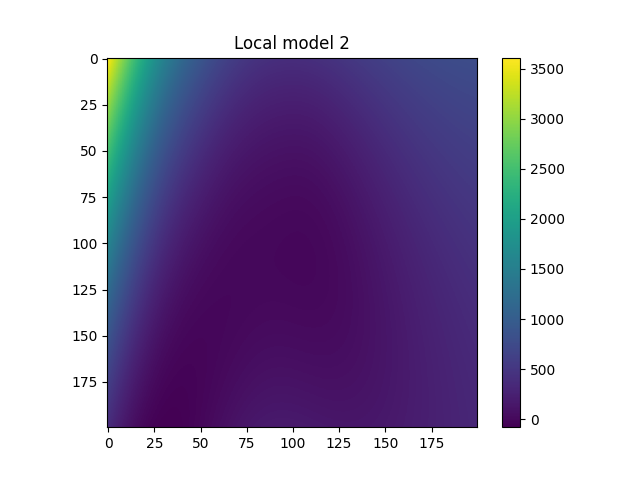
Plot local models#
for i in range(model.n_clusters):
plt.figure()
plt.imshow(
model.predict_local_model(fine_input, i)["rosen"].reshape((
refinement,
refinement,
))
)
plt.colorbar()
plt.title(f"Local model {i}")
plt.show()
Total running time of the script: (0 minutes 1.744 seconds)